Numpy, MATLAB and singular matrices
Last week I gave a live demo of the IPython notebook to a group of numerical analysts and one of the computations we attempted to do was to solve the following linear system using Numpy’s solve command.
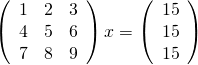
Now, the matrix shown above is singular and so we expect that we might have problems. Before looking at how Numpy deals with this computation, lets take a look at what happens if you ask MATLAB to do it
>> A=[1 2 3;4 5 6;7 8 9]; >> b=[15;15;15]; >> x=A\b Warning: Matrix is close to singular or badly scaled. Results may be inaccurate. RCOND = 1.541976e-18. x = -39.0000 63.0000 -24.0000
MATLAB gives us a warning that the input matrix is close to being singular (note that it didn’t actually recognize that it is singular) along with an estimate of the reciprocal of the condition number. It tells us that the results may be inaccurate and we’d do well to check. So, lets check:
>> A*x ans = 15.0000 15.0000 15.0000 >> norm(A*x-b) ans = 2.8422e-14
We seem to have dodged the bullet since, despite the singular nature of our matrix, MATLAB has able to find a valid solution. MATLAB was right to have warned us though…in other cases we might not have been so lucky.
Let’s see how Numpy deals with this using the IPython notebook:
In [1]: import numpy from numpy import array from numpy.linalg import solve A=array([[1,2,3],[4,5,6],[7,8,9]]) b=array([15,15,15]) solve(A,b) Out[1]: array([-39., 63., -24.])
It gave the same result as MATLAB [See note 1], presumably because it’s using the exact same LAPACK routine, but there was no warning of the singular nature of the matrix. During my demo, it was generally felt by everyone in the room that a warning should have been given, particularly when working in an interactive setting.
If you look at the documentation for Numpy’s solve command you’ll see that it is supposed to throw an exception when the matrix is singular but it clearly didn’t do so here. The exception is sometimes thrown though:
In [4]:
C=array([[1,1,1],[1,1,1],[1,1,1]])
x=solve(C,b)
---------------------------------------------------------------------------
LinAlgError Traceback (most recent call last)
in ()
1 C=array([[1,1,1],[1,1,1],[1,1,1]])
----> 2 x=solve(C,b)
C:\Python32\lib\site-packages\numpy\linalg\linalg.py in solve(a, b)
326 results = lapack_routine(n_eq, n_rhs, a, n_eq, pivots, b, n_eq, 0)
327 if results['info'] > 0:
--> 328 raise LinAlgError('Singular matrix')
329 if one_eq:
330 return wrap(b.ravel().astype(result_t))
LinAlgError: Singular matrix
It seems that Numpy is somehow checking for exact singularity but this will rarely be detected due to rounding errors. Those I’ve spoken to consider that MATLAB’s approach of estimating the condition number and warning when that is high would be better behavior since it alerts the user to the fact that the matrix is badly conditioned.
Thanks to Nick Higham and David Silvester for useful discussions regarding this post.
Notes
[1] – The results really are identical which you can see by rerunning the calculation after evaluating format long in MATLAB and numpy.set_printoptions(precision=15) in Python

I tried to get this result in Euler, using various methods, including the package ALgLib, which uses LAPACK routines, and the LAPACL routines of Maxima. Really, no success.
>A=redim(1:9,3,3); b=sum(A);
>A\b
Determinant zero!
Error in :
A\b
^
>alsolve(A,b,fit(A,b)
0
3
0
>svdsolve(A,b)
1
1
1
>&load(lapack);
>&dgesv(@A,@b)
[ 0.0 ]
[ ]
[ 3.0 ]
[ ]
[ 0.0 ]
>M=A|b; M=M/b
0.166667 0.333333 0.5 1
0.266667 0.333333 0.4 1
0.291667 0.333333 0.375 1
>pivotize(M,1,3)
0.333333 0.666667 1 2
0.133333 0.0666667 0 0.2
0.166667 0.0833333 0 0.25
>pivotize(M,2,1)
0 0.5 1 1.5
1 0.5 0 1.5
0 0 0 0
>
Sorry about the formatting. Cannot use ?
Mathematica will give the same solution and a warning, but no condition number.
LinearSolve[N@{{1,2,3}, {4,5,6}, {7,8,9}}, {15,15,15}]LinearSolve::luc: “Result for LinearSolve of badly conditioned matrix {{1.,2.,3.},{4.,5.,6.},{7.,8.,9.}} may contain significant numerical errors”
{-39., 63., -24.}
R will give no solution, just an error and a condition number.
solve(matrix(1:9, ncol=3, byrow=T), c(15,15,15))system is computationally singular: reciprocal condition number = 2.59052e-18
I’m not good with Maple and I could only get a symbolic (full) solution, even with inexact numbers.
Julia 0.1.2 gives the same solution, but no warning.
In my opinion, octave gives the best solution:
warning: matrix singular to machine precision, rcond = 3.08471e-18
warning: attempting to find minimum norm solution
warning: dgelsd: rank deficient 3×3 matrix, rank = 2
x =
-7.5000e+00
6.8782e-15
7.5000e+00
It is possible to get this solution by using the pseudo inverse of A
——————————————–
from numpy.linalg import pinv # python
x=dot(pinv(A),b)
——————————————–
x=pinv(A),b) % matlab
——————————————–
Nice blog
Thanks for all of the comments and evaluations in other systems.
@Szabolcs – As you probably discovered yourself, if you use Mathematica’s linear solve with exact arithmetic you get a completely different answer, which is interesting
In[1]:= LinearSolve[{{1, 2, 3}, {4, 5, 6}, {7, 8, 9}}, {15, 15, 15}]
Out[1]= {-15, 15, 0}
Maple doesn’t give a warning either but does give results compatible with most of the other systems:
with(LinearAlgebra):
M := Matrix([[1.0, 2.0, 3.0],[4.0, 5.0, 6.0],[7, 8, 9]]):
v := Vector([15, 15, 15]):
x:=LinearSolve(M,v):
lprint(x);
gives
Vector[column](3, {1 = HFloat(-38.9999999999999929), 2 = HFloat(62.9999999999999929), 3 = HFloat(-24.)}, datatype = float[8], storage = rectangular, order = Fortran_order, shape = [])
I’m relatively new to Maple myself so I am not yet sure how to just get the floating point numbers of the returned vector in a simple list like (a,b,c)
The Julia team are also looking at this issue now:
https://github.com/JuliaLang/julia/issues/4286
An issue was reported on NumPy but no response so far:
https://github.com/numpy/numpy/issues/3755
Good blog! A related question is how you can tell in this particular case that the matrix must be singular (i.e. must have a vanishing determinant) without explicitly computing its determinant.
Here’s something weird in MATLAB:
A = [1 2 3; 4 5 6; 7 8 9]
B = [1 4 7; 2 5 8; 3 6 9]
Up to now it looks like A’ == B and MATLAB claims it’s so. But now try
A’ \ [15;15;15]
Warning: Matrix is close to singular or badly scaled. Results may
be inaccurate. RCOND = 1.156482e-18.
ans = [-2.5; 0; 2.5]
B \ [15; 15; 15]
Warning: Matrix is singular to working precision.
ans = [NaN; NaN; NaN]
Does MATLAB not really carry out the transposition in this case, just change how it interprets the underlying data?
I realized that I entered the transpose of the matrix into Maple when I tried it. Interestingly, in this case it gives a symbolic solution, even if the matrix had inexact numbers:
Vector(3, {(1) = -5.+_t1[1], (2) = 5.-2*_t1[1], (3) = _t1[1]})
I also tried the transpose in other systems:
In MATLAB I get a warning, as before. (But see my other comment—it might return NaN.)
In Mathematica I get {-2.5, 0, 2.5} but no warning.
R gives no solution and says “Lapack routine dgesv: system is exactly singular: U[3,3] = 0”
Julia gives no result and says “ERROR: SingularException(3)”
What Szabolics is seeing is the effect of rounding errors.
It just so happens that LU factorization of B = A’ produces an exactly singular
U matrix, and this is what produces the NaNs in the solution.
But rounding errors make the U factor for A nonsingular:
>> B = zeros(3); B(:) = 1:9; A = B’; [LA,UA] = lu(A); [LB,UB] = lu(B); UA, UB
UA =
7.0000e+00 8.0000e+00 9.0000e+00
0 8.5714e-01 1.7143e+00
0 0 1.1102e-16
UB =
3 6 9
0 2 4
0 0 0
Additional background to supplement Nick’s answer to Szabolcs:
The A’\[15;15;15] case goes through an optimized code path that avoids doing an explicit transpose. The different results are then different round-off behaviors from two separate code paths.
@Szabolcs:
In your example, A’\b should be similar to doing: linsolve(A,b,struct(‘TRANSA’,true)). This selects a separate execution path specifically to solve a system of the form A’*x=b, which explains the different results.
Of course mldivide will first perform tests on the matrix A to check for special properties and chose an appropriate solver [1], while linsolve relies on the user to specify the properties of the coefficient matrix..
The underlying LAPACK routine used to solve [2] a general double-precision real matrix is DGESV. The expert version of this driver (DGESVX) let us specify whether to solve for Ax=b or A’x=b (second parameter to the function) [3].
[1]: http://www.mathworks.com/support/solutions/en/data/1-172BD/
[2]: http://www.netlib.org/lapack/lug/node26.html
[3]: http://www.netlib.org/lapack/explore-html/dgesvx.f.html
Note that my comment above only holds when using the operator form A’\b not the function form mldivide(A’,b) of the backslash.
My guess is that the MATLAB parser specifically recognizes the first case and optimizes it accordingly at an early stage, but not the second form where the matrix would already be transposed before handing it to mldivide…
Is there a function in IPython or numpy already to get the estimate of the matrix conditional or its reciprocal?
Warren
There’s numpy.linalg.cond but, long term, there’s a better way to find the estimate in numpy’s solve. As it says within the discussion at https://github.com/numpy/numpy/issues/3755 solve currently uses LAPACK’s [DZ]GESV but if they used [DZ]GESVX instead they’d get the reciprocal condition number along with the solution.
Mike
This is timely. We are about to hit linear and non-linear equations in my “computational methods in physics” course, and I can use this as an example of of being aware of what might go wrong. I was showing them how to get a wrong answer out of quad (using scipy.integrate) today.
Sounds like an interesting topic for a blog post, Jim
@Mike Croucher
Go right ahead. Here is simple code to calculate the same integral twice and get one right and one wrong result. Explaining why was a homework problem.
from scipy.integrate import quad
from scipy import exp
from numpy import inf
def bb(x):
return x**3/(exp(x)-1)
print quad(bb,0,inf)
print ‘\n We can fool it.’
print ‘Here we do the same calculation with a change of variable’
def bb2(t):
x=1e5*t
return 1e5*(x**3/(exp(x)-1))
print quad(bb2,0,inf)
@Mike Croucher
Today, we have done that and it will be available in SciPy version 0.19. See the dev version of the documentation.
Hopefully soon we will see some action on the NumPy side too.