A portable Jupyter (IPython) install on Windows with an R kernel
I recently found myself in need of a portable install of the Jupyter notebook which made use of a portable install of R as the compute kernel. When you work in institutions that have locked-down managed Windows desktops, such portable installs can be a life-saver! This is particularly true when you are working with rapidly developing projects such as Jupyter and IRKernel.
It’s not perfect but it works for the fairly modest requirements I had for it. Here are the steps I took to get it working.
Download and install Portable Python
I downloaded Portable Python 2.7.6.1 from http://portablepython.com/ and installed into a directory called Portable Python 2.7.6.1
Update IPython and install the extra modules we need
This version of Portable Python comes with a portable IPython instance but it is too old to support alternative kernels. As such, we need to install a newer version.
Open a cmd.exe command prompt and navigate to Portable Python 2.7.6.1\App\Scripts.
Enter the command
easy_install ipython.exe
You’ll now find that you can launch the ipython.exe terminal from within this directory:
C:\Users\walkingrandomly\Desktop\Portable Python 2.7.6.1\App\Scripts>ipython Python 2.7.6 (default, Nov 10 2013, 19:24:18) [MSC v.1500 32 bit (Intel)] Type "copyright", "credits" or "license" for more information. IPython 3.1.0 -- An enhanced Interactive Python. ? -> Introduction and overview of IPython's features. %quickref -> Quick reference. help -> Python's own help system. object? -> Details about 'object', use 'object??' for extra details. In [1]: exit()
If you try to launch the notebook, however, you’ll get error messages. This is because we haven’t taken care of all the dependencies. Let’s do that now. Ensuring you are still in the Portable Python 2.7.6.1\App\Scripts folder, execute the following commands.
easy_install pyzmq easy_install jinja2 easy_install tornado easy_install jsonschema
You should now be able to launch the notebook using
ipython notebook
Install portable R and IRKernel
- I downloaded Portable R 3.2 from http://sourceforge.net/projects/rportable/files/ and installed into a directory called R-Portable
- Move this directory into the Portable Python directory. It needs to go inside Portable Python 2.7.6.1\App (see this discussion to learn how I discovered that this location was the correct one)
- Launch the Portable R executable which should be at Portable Python 2.7.6.1\App\R-Portable\R-portable.exe and install the IRKernel packages by doing
install.packages(c("rzmq","repr","IRkernel","IRdisplay"), repos="http://irkernel.github.io/")
Install additional R packages
The version of Portable R I used didn’t include various necessary packages. Here’s how I fixed that.
- Launch the Portable R executable which should be at Portable Python 2.7.6.1\App\R-Portable\R-portable.exe and install the following packages
install.packages('digest') install.packages('uuid') install.packages('base64enc') install.packages('evaluate') install.packages('jsonlite')
Install the R kernel file
Create the directory structure Portable Python 2.7.6.1\App\share\jupyter\kernels\R_kernel
Create a file called kernel.json that contains the following
{"argv": ["R-Portable/App/R-Portable/bin/i386/R.exe","-e","IRkernel::main()",
"--args","{connection_file}"],
"display_name":"Portable R"
}
This file needs to go in the R_kernel directory created earlier. Note that the kernel location specified in kernel.json uses Linux style forward slashes in the path rather than the backslashes that Windows users are used to. I found that this was necessary for the kernel to work –it was ignored by the notebook otherwise.
Finishing off
Everything created so far, including R, is in the folder Portable Python 2.7.6
I created a folder called PortableJupyter and put the Portable Python 2.7.6 folder inside it. I also created the folder PortableJupyter\notebooks to allow me to carry my notebooks around with the software that runs them.
There is a bug in Portable Python 2.7.6.1 relating to scripts like IPython.exe that have been installed using easy_install. In short, they stop working if you move the directory they’re installed in – breaking portability somewhat! (Details here)
The workaround is to launch Ipython by running the script Portable Python 2.7.6.1\App\Scripts\ipython-script.py
I didn’t want to bother with that so created a shortcut in my PortableJupyter folder called Launch notebook. The target of this shortcut was the following line
%windir%\system32\cmd.exe /c "cd notebooks && "%CD%/Portable Python 2.7.6.1/App\python.exe" "%CD%/Portable Python 2.7.6.1\App\Scripts\ipython-script.py" notebook"
This starts the notebook using the default web browser and puts you in the notebooks directory.
The pay off
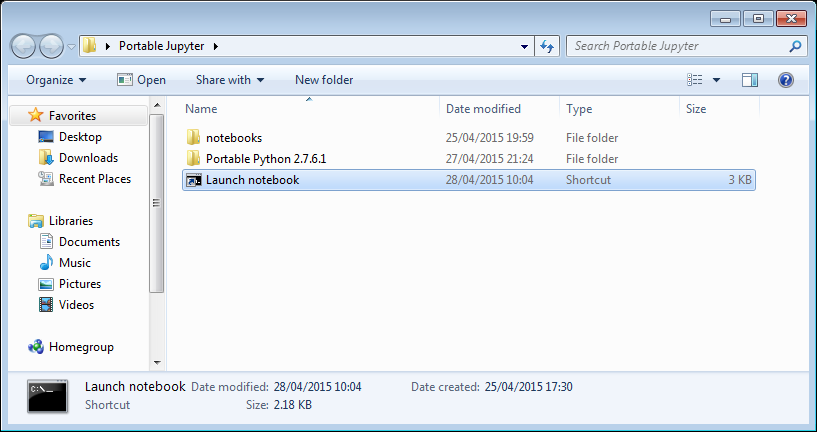
My folder looks like this:
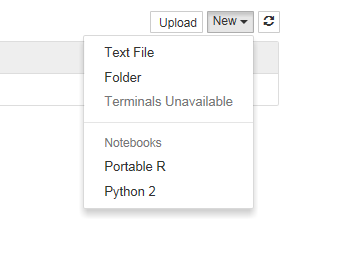
If I click on the Launch Notebook shortcut, I get a Jupyter session with 2 kernel options
I can choose the Portable R kernel and start using R in the notebook!


Nice. Did you know that the conda package repository, binstar, now has an R channel? You should be able to most of the above with the following after installing miniconda and setting your path:
conda install -c r ipython-notebook r-irkernel
That’s fantastic! Can I install completely portable versions of Python/R using conda? I.e. ones that don’t need admin rights on Windows to install and don’t leave any residue behind? That was the driving factor in going down this route.
For integrating with our managed desktop ‘properly’, however, I intend to use Anaconda. The fact that I can add R-support so easily now that you’ve given me the above information makes it even easier :)
Yes if you’re using Linux; haven’t tried doing anything similar on Windows but imagine it would work.
A follow-up post on setting up jupyterhub for students at the University of Sheffield would be interesting. There have been several posts on the ipython-dev list in recent months on people using jupyter for different courses; I’m sure the ipython-dev team would be interested in hearing about anything you set up for students. FYI Jessica Hamrick’s post on the Rackspace dev blog gives a particularly good overview of the current usability of the project for this purpose.
Windows is vital for teaching – at Sheffield at least.
I’ll use the portable install created above to get us through the next two weeks which we are using as a dry run and pre-requisite set of sessions for a Bioinformatics module which will be delivered in October.
I’ll work with the managed desktop team at Sheffield to get the Anaconda solution ‘packaged’ by then — kernels in Python 2, Python 3 and R to start with – all running on the Windows desktop.
Longer term, I’d like to do something more interesting with this technology and the blogpost you mentioned (this one, yes? https://developer.rackspace.com/blog/how-did-we-serve-more-than-20000-ipython-notebooks-for-nature/ ) looks like the kind of thing that would be fun to do :)
Nope, it was https://developer.rackspace.com/blog/deploying-jupyterhub-for-education/. The jupyterhub demo for Nature was concerned with providing Notebooks to anonymous users whereas Jessica was more interested in students being able to authenticate against the notebook server, complete Notebook-based coursework then the tutor grade submissions manually or automatically using a unit testing approach (see https://github.com/jupyter/nbgrader).
Are there plans to introduce researchers to Jupyter using something like Software Carpentry workshops? This would be awesome, particularly if researchers could ‘try before they buy’ by being able to participate in the course using their own machines without having to install anything locally.
There are plans but so far just a twinkle in my eye. I hope to have more time to devote to it soon though.
@Will Furnass
This is awesome!
Thank Will Furnass. The tip is awesome!
Hello!
I created virtual environment using conda in folder “dev”. Environment folder is named “pyenv”. There jupyter notebook, numpy and matplotlib in it. I created shortcut file in it. In “dev” folder I created .lnk file and typed %windir%\system32\cmd.exe /c “”%CD%\pyenv\python.exe” “%CD%\pyenv\Scripts\ipython-script.py” “notebook”. After doubleclick I see console with message “ipython notebook is deprecated will be removed in future versions. You likely want to juse jupyter notebook.” After 5 seconds jupyter notebook starts.
I decided to modify relative link in .lnk file. New version is %windir%\system32\cmd.exe /c “”%CD%\pyenv\python.exe” “%CD%\pyenv\Scripts\jupyter-script.py” “notebook”. But at this case Jupyter notebook doesn’t start.
Then I noticed strange: when I open cmd in “dev” folder and type in console pyenv\Scripts\ipython.exe notebook – all works fine. But when I type pyenv\Scripts\jupyter.exe notebook – I see error, that notebook isn’t jupyter command.
How can I fix it?
Best regards, Andrey.
I found answer myself. For Jupyter Notebook in shortcut properties target should be %windir%\system32\cmd.exe /c “%CD%\pyenv\Scripts\jupyter-notebook.exe” and start in should be empty.
Thanks so much Will!
I got it working with PortableR and Anaconda3.
The issue I had was to get jupyter in anaconda3 to “detect” the path for the R kernel.
Turns out the solution was to add the `Rkernel/kernel.json` in the directory holding `python3/kernel.json` in anaconda3.